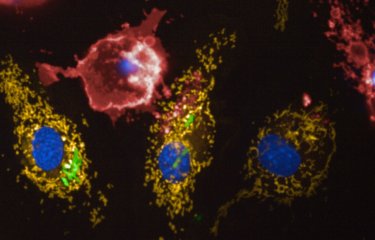
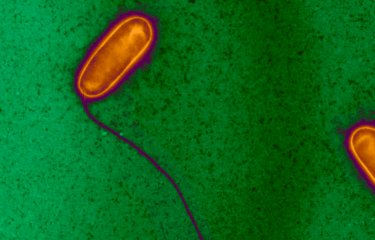
Legionellosis or Legionnaire's disease affected more than 1,000 people in France in 2003 and caused nearly 130 deaths. This emerging disease is caused by Legionella pneumophila, an environmental bacterium that can grow in hot water systems. A team at the Institut Pasteur associated with the CNRS [French National Center for Scientific Research] and in collaboration with the National Reference Center for Legionella, Inserm [National Institute for Health and Medical Research] in Lyon, compared the genome sequence of the strain responsible for an epidemic that recently occurred near Lens (Nord-Pas-de-Calais region) with that of an endemic strain frequently isolated in France. This work, published in Nature Genetics, may allow now to better understand how certain Legionella strains are spread in the population by aerosols originating from cooling towers. These results should also help to identify new targets for the development of tools to fight these bacteria.
Press release
Paris, october 4, 2004
The name Legionnaire’s disease was coined due to an epidemic of legionellosis that occurred in 1976 where Legionella pneumophila affected 200 participants of the 58th American Legion Convention in Philadelphia. Since then, many epidemics have been observed in North America and Europe. An estimated 8,000 to 18,000 people die of Legionnaire’s disease each year in the United States. In France, although sporadic cases are regularly reported throughout the country, eight particularly serious epidemics originating from cooling towers have also occurred since 1998. Legionella are parasites of amoebae present in fresh water and artificial water systems, but are also able to cause disease in man, through respiratory passages, once they are spread in the air via aerosols.
The team led by Carmen Buchrieser of the laboratory of Genomics of Microbial Pathogens at Institut Pasteur, (directed by Frank Kunst and Philippe Glaser) in association with the CNRS (1) and in collaboration with the Génopole® Pasteur and the National Reference Center for Legionella (directed by Jérôme Etienne (2)), sequenced and compared the genomes of two Legionella strains: one isolated from a nosocomial epidemic that had occurred at Hôpital Georges Pompidou (Paris) in 2000 (named strain "Paris"), and the other one responsible for a serious epidemic (86 cases and 17 deaths) that occurred in the North of France in winter 2003-2004 (named strain "Lens"). The researchers aimed to better characterize the genetic basis of the virulence of Legionella pneumophila and to understand differences occurring between these strains. Both belong to the same serogroup, seropgroup 1, that is alone responsible for 84% of reported cases of Legionnaire’s disease.
Carmen Buchrieser and her colleagues have proven that Legionella pneumophila are subject to significant genome variations, since about 13% differences were observed between the two strains studied. They identified a large number of genes coding proteins that might contribute to the adaptation of the bacteria to humans, like proteins similar to those of superior organisms (amoebae to man), others possessing domains probably capable of modifying the physiology of host cells, which might be potential virulence factors. The identification of several hundred genes that differ among the "Paris" and the "Lens" strain together with its analysis should make it possible to eventually understand what made certain strains particularly virulent and enabled them to spread so dramatically through the population.
This work sheds light on the evolution of Legionella pneumophila over time and should make it possible to understand how this bacterium is capable of adapting to hosts as different as amoebae (microorganisms living in aqueous environments) and man. These new data also open new avenues to create improved diagnostics as well as for the development of new, effective therapeutic weapons and biocides for decontaminating water systems.
______________________________________________________
1 Unité Génétique des génomes [Genome Genetics Unit], CNRS
2 Inserm E-0230, Faculty of Medecine, Lyon
Sources
Evidence in the Legionella pneumophila genome for exploitation of host cell functions and high genome plasticity. Nature Genetics, November 2004 (vol 36 :1165-1173).
Christel Cazalet* (1), Christophe Rusniok (1), Holger Brüggemann, Nora Zidane (2), Arnaud Magnier (2), Laurence Ma (2), Magalie Tichit (2), Sophie Jarraud (3), Christiane Bouchier (2), François Vandenesch (3), Frank Kunst (1), Jerome Etienne (3), Philippe Glaser (1), and Carmen Buchrieser (1)
(1) Pathogenic Microorganism Genomics Laboratory, CNRS URA 2171, Institut Pasteur
(2) Genomics Platform, Pasteur Génopole Ile de France, Institut Pasteur
(3) National Legionella Reference Center, Bacteriology Laboratory INSERM E-
0230, Faculty of Medecine, IFR 62, Lyon
* CNRS Scholarship - Veolia Water - Anjou Recherche
Contact persons
Institut Pasteur- Press Office
Nadine Peyrolo
01 44 38 91 30 - npeyrolo@pasteur.fr
Bruno Baron
01 44 38 91 30 - bbaron@pasteur.fr
CNRS - Press Office
Isabelle Tratner
01 44 96 49 88 - isabelle.tratner@cnrs-dir.fr
Inserm - Press Office
Martine Moëllic
01 44 23 60 97 - presse@tolbiac.inserm.fr