A study by the Institut Pasteur has highlighted the genetic diversity of Salmonella enterica serovar Panama, a bacterial pathogen responsible for frequent foodborne infections in the French West Indies and French Guiana, and revealed the worrying evolution of a bacterium that was previously susceptible to treatment. The findings emphasize the need for increased surveillance to prevent health risks. In early 2026, the Le Roch-Les Mousquetaires Foundation renewed its support for the work of François-Xavier Weill's team in monitoring the evolution of bacterial resistance to antibiotics.
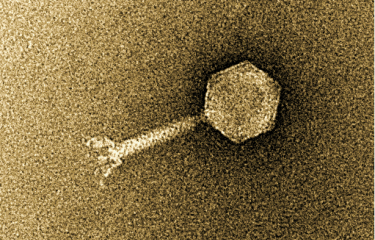
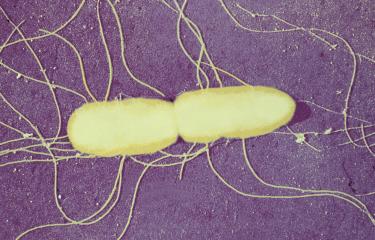
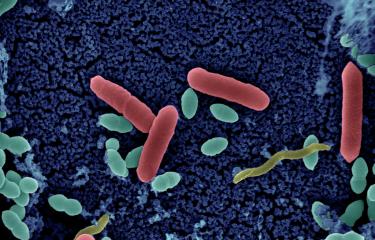
Salmonella enterica serovar Panama, one of the most common causative agents of non-typhoidal Salmonella disease in France's overseas territories, is surprisingly diverse. An unprecedented phylogenetic study led by Professors François-Xavier Weill (Institut Pasteur) and Kate Baker (University of Cambridge) and published in 2025 in The Lancet Microbe analyzed more than 800 genomes collected in 45 countries since 1931. This is the most extensive analysis ever carried out on S. Panama, which caused significant hospital outbreaks in mainland France in the 1970s, with more than a thousand cases in some years.
Clades with contrasting profiles
The scientists identified two clades, mainly found in Europe and Asia, that are particularly resistant to antibiotics and more invasive. The clades circulating in the Americas, including the French West Indies, remain susceptible to treatment. These differences can be explained by different reservoirs:
- In Europe, the resistant clade is thought to be linked to the development of the pig farming industry after the Second World War.
- In Asia, some resistant strains could come from complex food supply chains.
- In the Americas, the strains have remained susceptible to antibiotics because they are from an environmental reservoir (maybe reptiles). These strains may have been introduced into the French West Indies as a result of population movements linked to the construction of the Panama Canal.
A health risk that needs monitoring
"These results show that Salmonella enterica serovar Panama is not a uniform pathogen but rather a series of clades that behave in different ways," explains François-Xavier Weill. The presence of invasive strains with multiple antibiotic resistance in Europe and Asia confirms that more vigilance is needed, especially in food supply chains.
A project supported by the Le Roch-Les Mousquetaires FoundationThis research is part of a broader project, funded for the past few years by the Le Roch-Les Mousquetaires Foundation, aimed at shedding light on the development of bacterial resistance. "Their support is crucial in helping us elucidate these mechanisms and propose effective prevention strategies," continues the scientist. At the beginning of 2026, the foundation renewed its support until 2028. |
Source : Global diversity and evolution of Salmonella enterica serovar Panama: a genomic epidemiology study, The Lancet Microbe, September 2025
Caisey V Pulford1, Blanca M Perez-Sepulveda1, Danielle J Ingle2, Rebecca J Bengtsson1, Rebecca J Bennett1, Ella V Rodwell3, Maria Pardos de la Gandara4, Charlotte E Chong5, P Malaka De Silva5, Magali Ravel4, Véronique Guibert4, Elisabeth Njamkepo4, Neil Hall6, Marie A Chattaway3, Benjamin P Howden7, Deborah A Williamson8, Jay C D Hinton1, François-Xavier Weill9, Kate S Baker10
1Clinical Infection, Microbiology, and Immunology, Institute of Infection, Veterinary and Ecological Sciences, University of Liverpool, Liverpool, UK.
2Department of Microbiology and Immunology, The University of Melbourne at The Peter Doherty Institute for Infection and Immunity, Melbourne, VIC, Australia.
3Gastrointestinal Bacteria Reference Unit, UK Health Security Agency, London, UK.
4Unité des Bactéries pathogènes entériques, Institut Pasteur, Université Paris Cité, Paris, France.
5Department of Genetics, University of Cambridge, Cambridge, UK.
6Earlham Institute, Norwich Research Park, Norwich, UK.
7Microbiological Diagnostic Unit Public Health Laboratory, Department of Microbiology and Immunology, The University of Melbourne at The Peter Doherty Institute for Infection and Immunity, Melbourne, VIC, Australia; Centre for Pathogen Genomics, The University of Melbourne, Melbourne, Australia.
8School of Medicine, University of St Andrews, Fife, Scotland.
9Unité des Bactéries pathogènes entériques, Institut Pasteur, Université Paris Cité, Paris, France.
10Clinical Infection, Microbiology, and Immunology, Institute of Infection, Veterinary and Ecological Sciences, University of Liverpool, Liverpool, UK; Department of Genetics, University of Cambridge, Cambridge, UK.