Scientists from the Institut Pasteur and the CNRS, working in collaboration with US and Australian teams, have completed full genomic and evolutionary characterization of the strain of the Elizabethkingia anophelis bacterium that affected around 60 people in Wisconsin in 2015-2016. Their research reveals that this outbreak was caused by a highly mutant strain of E. anophelis. It is unusual for bacteria to have such a high rate of mutation, and this property could confer a selective advantage on this strain – which is also notable for its resistance to several antibiotics. These data, published on May 24, 2017 in the journal Nature Communications, have been made available to the international scientific community.
In November 2015, an outbreak caused by the little-known bacterium Elizabethkingia anophelis was reported in Wisconsin, in the United States. A total of 66 cases were confirmed in the country – an exceptional number for an outbreak caused by this bacterium. Despite the introduction of an antibiotic treatment protocol, the outbreak claimed the lives of 21 patients, most of whom were over the age of 65 and had at least one serious underlying condition.
To shed light on this unusual outbreak, scientists from the Microbial Evolutionary Genomics Unit (Institut Pasteur/CNRS), the French Institute of Bioinformatics (CNRS/Inserm/Inra/Inria/CEA), and the Bioinformatics, Biostatistics and Integrative Biology (Institut Pasteur/CNRS,) led at the Institut Pasteur by Sylvain Brisse, in collaboration with teams from the Centers for Disease Control and Prevention (CDC, USA), the Wisconsin Department of Health Services, the Wisconsin State Laboratory of Hygiene and the University of Melbourne (Australia), carried out extensive investigative genomic research on isolates from patients affected by this outbreak.
The French scientists had previously described several strains of this bacterium during an infection episode in the Central African Republic and compared the data to genomes in public sequence databases. This research enabled them to define a standardized typing system, which they also used to characterize the strain behind the Wisconsin outbreak.
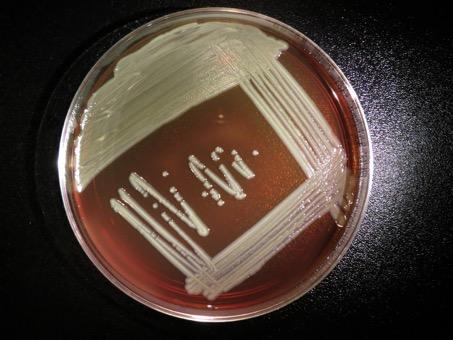 |
Elizabethkingia anophelis is a bacterium that was first isolated from Anopheles mosquitoes and described in 2011. But there is no indication that the mosquito plays a role in transmitting the bacterium to humans, and it seems more likely that environmental contamination, for example through water, may be responsible. It is very rare for Elizabethkingia bacteria to cause infections in humans, but they are capable of colonizing the respiratory system. In the event of infection, symptoms may include fever, shortness of breath and chills. E. anophelis was previously responsible for two cases of neonatal meningitis in the Central African Republic and a small outbreak involving six cases of infection in an intensive care unit in Singapore. © CDC special bacteriology reference lab.© CDC special bacteriology reference lab. |
Sequencing of the bacterial isolates, performed by the Atlanta-based CDC, firstly revealed that the outbreak was "clonal"; in other words it was caused by a single strain of E. anophelis. All the cases of infection associated with the outbreak therefore represented a new lineage of E. anophelis bacteria. It should be noted that this was a community outbreak, which is unusual for Elizabethkingia, since previous infections have always occurred within healthcare facilities.
But the most striking finding was the highly mutant nature of the strain responsible for the outbreak. "We demonstrated that the strain had undergone nearly 300 mutations in just one year – ten times more than 'traditional' bacteria. This high mutation rate is extremely rare for bacterial outbreaks," comments Sylvain Brisse. The scientists demonstrated that this characteristic could be explained by the insertion of a mobile genetic element in a DNA repair gene within the bacterial genome.
Although the many accumulated mutations did not directly make the strain more resistant to antibiotics – E. anophelis is already known to be highly resistant –, the scientists believe that the high mutation rate has given the bacterium a selective advantage, facilitating its survival in the environmental reservoir or the infectious source. This property may also have increased its pathogenicity in humans. The scientists also identified other specific characteristics including detoxification genes or genes for the utilization of organic plant matter. The latter might have helped the bacteria to colonize the environmental reservoir more effectively, which may partly explain the scale of the outbreak.
These observations prompted the scientists to trace the diversification of the bacteria during the outbreak. They dated the emergence of their common ancestral strain to one year before the first confirmed case of infection. "This information provided an additional temporal dimension to our knowledge of the outbreak, enabling us to develop new epidemiological theories and potentially leading to the retrospective identification of new cases," continues the scientist.
This study not only led to the discovery of a bacterial mutant responsible for an outbreak; it also gave the scientists the opportunity to highlight the importance of real-time data exchange during outbreaks – in line with the ideals of the "open data" movement – as a means of providing the best possible response to public health emergencies.
This research was funded by the Institut Pasteur, the French National Center for Scientific Research, the French Foundation for Medical Research, the French Institute of Bioinformatics, the Integrative Biology of Emerging Infectious Diseases Laboratory of Excellence (IBEID LabEx, managed by the French National Research Agency), and the Advanced Molecular Detection (AMD) Initiative led by the CDC (Atlanta, USA).
Source
Evolutionary dynamics and genomic features of the Elizabethkingia anophelis 2015 to 2016 Wisconsin outbreak strain, Nature Communications, 24 mai 2017.
Amandine Perrin (a,b,c)&, Elise Larsonneur (a,b,d)&, Ainsley C. Nicholson (e)&, David J. Edwards (f,g), Kristin M. Gundlach (h), Anne M. Whitney (e), Christopher A. Gulvik (e), Melissa E. Bell (e), Olaya Rendueles (a,b), Jean Cury (a,b), Perrine Hugon (a,b), Dominique Clermont (i), Vincent Enouf(j), Vladimir Loparev (k), Phalasy Juieng (k), Timothy Monson (h), David Warshauer (h), Lina I Elbadawi (l, m), Maroya Spalding Walters (n), Matthew B Crist (n), Judith Noble-Wang (n), Gwen Borlaug (m), Eduardo P.C. Rocha (a,b), Alexis Criscuolo (c,j), Marie Touchon (a,b), Jeffrey P. Davis (m), Kathryn E. Holt (f,g), John R. McQuiston (e),*, Sylvain Brisse (a,b,o),*
- Institut Pasteur, Microbial Evolutionary Genomics, F-75724, Paris, France
- CNRS, UMR 3525, F-75724, Paris, France
- Institut Pasteur, Bioinformatics and Biostatistics HUB, C3BI, USR 3756 IP CNRS, F-75724, Paris, France
- CNRS, UMS 3601 IFB-Core, F- 91198, Gif-sur-Yvette, France
- Special Bacteriology Reference Laboratory, Bacterial Special Pathogens Branch, Division of High Consequence Pathogens and Pathology, Centers for Disease Control and Prevention, Atlanta, Georgia 30329, USA
- Centre for Systems Genomics, University of Melbourne, Parkville, Victoria 3010, Australia
- Department of Biochemistry and Molecular Biology, Bio21 Molecular Science and Biotechnology Institute, University of Melbourne, Parkville, Victoria 3010, Australia
- Wisconsin State Laboratory of Hygiene, Madison, Wisconsin 53718, USA
- CIP – Institut Pasteur Collection, Institut Pasteur, F-75724, Paris, France
- Institut Pasteur, Pasteur International Bioresources Network (PIBnet), Mutualized Platform for Microbiology (P2M), F-75724, Paris, France
- Division of Scientific Resources, Centers for Disease Control and Prevention, Atlanta, Georgia 30329, USA
- Epidemic Intelligence Service, Centers for Disease Control and Prevention, Atlanta, Georgia 30329, USA
- Division of Public Health, Wisconsin Department of Health Services, Madison, Wisconsin 53701, USA
- Division of Healthcare Quality Promotion, Centers for Disease Control and Prevention, Atlanta, Georgia 30329, USA
- Institut Pasteur, Molecular Prevention and Therapy of Human Diseases, F-75724, Paris, France
& These authors contributed equally to this work.
* Authors to whom correspondence about the article should be addressed





