Identifying risk factors for disease, helping diagnose cancer, guiding therapeutic choices and monitoring the efficacy of treatment, facilitating surveillance of pathogens – in a huge number of fields, technological progress is now providing biomedical research with vast reams of information (or "big data") about the diseases under study, representing an extraordinary source of new knowledge and medical progress. But the human mind can no longer deal with these data flows by itself – it needs the help of increasingly complex computer tools to manage all this information. So how exactly does it all work?
Genomics: unleashing the potential of big data
Let's take the example of genomics, one of the fields that has become synonymous with the emergence of big data. Obtaining the first sequence of the human genome (our 23,000 genes, representing a "text" of 3 billion letters), which was finally revealed in 2003, took nearly 15 years and cost almost $3 billion. Nowadays we can obtain the sequence of an individual's genome within a day, using "high-throughput" sequencing techniques, for less than $1,000. This low cost – which is continuing to fall – means that the genomes of patient volunteers can be sequenced in any number of studies on conditions such as autism, cancer and neurodegenerative diseases. Not long ago, geneticists based their research on the analysis of a few genes in a limited number of patients. Today they can access information about the entire genome of hundreds or even thousands of people, which they can associate with other data about these patients (age, gender, existing conditions or medication, etc.). The major challenge for today's researchers is how to search and analyze these data, which are stored in "digital warehouses" – huge databases that can be consulted online.

Making sense of data
This can best be illustrated by a specific example: let's say that we want to compare dozens of genomes of patients responding to treatment with those of patients in whom the same treatment is ineffective, to see whether there are any genetic differences between the two groups so that we can identify markers that will predict the efficacy of the therapy. A study of this nature is not "humanly" possible: we need a computer tool, which, once it has been programmed, will carry out this large-scale comparison, review the 3 billion letters in each genome, identify differences in the sequences of the genomes and pinpoint those that can be linked to the response to treatment. A significant part of the work of the biologists leading the study involves liaising with bioinformaticians, statisticians and other mathematicians to design and develop IT and statistical programs and algorithms for their study. Because, as all scientists point out, "data on their own are pointless – we need to give them meaning."

The exponential production of genome sequences is not restricted to the human genome. Scientists are busy sequencing all the DNA in samples of patients' stools (this is referred to as metagenomics) to study the gut microbiota – the 100,000 microbial species in our gut flora whose composition varies from one individual to the next and which have a major influence on our health –, or the genomes of viruses, bacteria and other microbes that threaten us, so that we are better placed to monitor them (see below inset on PIBnet).
Monitoring pathogens more effectively
The Institut Pasteur houses 14 National Reference Centers (CNRs), in association with Santé Publique France, which are responsible for monitoring infectious diseases in France and raising the alert in the event of an outbreak such as influenza, foodborne infection (salmonellosis, listeriosis, etc.), rabies or bacterial meningitis. "For the past two years, as part of the PIBnet* program, the genomes of strains of bacteria, viruses and fungi isolated in patients in France and sent to be analyzed at several of these centers – currently around 25,000 strains a year – are systematically sequenced on a shared platform, and all the data about each strain (provenance, virulence, any presence of resistance genes, etc.) are stored," explains Sylvain Brisse, Head of the Biodiversity and Epidemiology of Bacterial Pathogens Unit at the Institut Pasteur and holder of a Chair of Excellence for Genomic Epidemiology in Global Health. "This database enables us to compare our information with that obtained in other countries so that we can monitor the spread of pathogens at global level."
* Pasteur International Bioresources network.
A tsunami of images

The second field that has been revolutionized by the big data era is imaging, whether medical imaging or microscopy. Increasingly powerful and automated microscopes are unleashing a tsunami of images: in just an hour, the Titan Krios™, the world's most powerful microscope, can provide a quantity of images equivalent to 500 high-definition films (see below), which would be impossible for human brains to analyze within a reasonable time frame. Once again, the development of ad hoc computer tools is necessary. When it comes to image analysis, researchers are making increasing use of artificial intelligence, especially deep learning techniques, which have emerged in response to the growth of big data and the increase in computing power (see Interview, below). Last year, using a hundred thousand images of moles that had already been characterized as benign or malignant, a US team was able to teach a computer to diagnose melanomas. A recent study compared the machine's performance to that of 58 specialist physicians from 17 countries. The researchers noted that "most of the dermatologists performed less well," with an average of 87% of correct diagnoses as opposed to 95% for the computer. When well "trained", the computer is therefore more effective – and does not suffer from fatigue or lapses in concentration! These techniques are currently revolutionizing disease diagnosis, and the interpretation of medical images (obtained from X-rays, CT and MRI scans, microscopic observation of biopsy tissue, etc.) will increasingly become a task for computers, which are faster and more reliable, giving physicians more time to spend on other research activities and on doctor-patient dialog.

Big data is a key driver in the current revival of artificial intelligence
You direct a team that specializes in imaging.* How has big data changed things?
Together with the increase in computing power, big data has brought up to date some fairly old techniques in the field of artificial intelligence, which are now providing results precisely because we have access to big data. One particular approach is "deep learning", which no one was talking about just 10 years ago. With these techniques, we can now train an algorithm to distinguish between images of dogs and cats, for example, by giving it thousands of images to learn from (annotated with 0 for dogs and 1 for cats). After performing a huge number of calculations on these training data, the algorithm becomes able to generalize and recognize a cat or a dog from an unknown image. The more data we enter, the fewer mistakes it makes. The voice recognition on your smartphone, online machine translation services, self-driving cars and robots that can handle objects all draw on deep learning, which is now also arriving in the field of medicine. Although it is starting to be used in genomics, medical imaging is undoubtedly the area in which it is currently having the greatest impact.
What are the prospects for medical applications of artificial intelligence?
We are working with the Experimental Neuropathology Unit, directed by Fabrice Chrétien, on diagnosing cancerous brain tumors in children using post-operative biopsies from the Sainte-Anne and Necker Hospitals in Paris. Currently, for a given biopsy, two pathologists may disagree on the diagnosis in a significant proportion of cases. Being able to establish a highly precise diagnosis of the type of tumor is crucial in deciding on the course of treatment: chemotherapy, radiotherapy, additional surgery, etc. To analyze the biopsy, the expert has an image that typically contains 5 billion pixels (by way of comparison, a standard photo is composed of roughly 1 million pixels) and has no time to fully inspect it at its maximum resolution. As with the example of cats and dogs, we are teaching an algorithm to determine the degree of severity of a tumor by training it using hundreds of images of biopsies that have been correctly characterized by a group of experts. Our long-term objective is to facilitate and complement the histopathological analysis performed by the human expert, to make it more reliable but also faster, since it can take several days for a specialist to have access to all the tests needed to analyze a biopsy. In the meantime, before we achieve that objective, the tools that we are developing could already be used to draw the expert's attention to areas of the image that it may be worth studying in more detail.
Could these methods also speed up research?
Absolutely. At the Institut Pasteur, the team led by Arnaud Echard (in the Membrane Traffic and Cell Division Unit) is studying the final stage in cell division, abscission, which results in cell separation. When this process goes wrong, it can lead to the formation of many types of cancer. Arnaud's team identified 400 genes that are potentially involved in this process. To determine the exact role of each of these genes, they need to be inactivated one by one, then the cell divisions need to be filmed to determine the exact moment of separation, which varies from one cell to the next. Scientists carry out this task by inspecting the films with the naked eye, but to identify the impact of a single gene they need to observe roughly 300 cell divisions, representing 15 hours of work – which needs to be done three times to make sure it is reproducible, making a total of 45 hours per gene. If we multiply this by 400, it comes to 18,000 hours, or several years of full-time work! So we are introducing a deep learning algorithm with the aim of analyzing each film almost instantaneously. If successful, this technique will tell the team which genes influence abscission virtually straight away, potentially paving the way for the development of cancer treatment.
*Christophe Zimmer trained as an astrophysicist. His team is composed of mathematicians, physicists, computer scientists, biologists and one chemist.
Titanic quantities of data!
In July 2018, the Institut Pasteur officially opened its Nocard building, which houses the world's most powerful microscope, the Titan Krios™ – the only one of its kind in France used for health research –, acquired with the support of several donors. This fully automated cryo-electron microscope enables atomic-scale observation. It is also capable of examining several samples at any one time, and is therefore a huge generator of big data. "Storing the raw data generated by the Titan is a first major challenge – it could easily fill up all the hard disks on campus in no time!" explains Michael Nilges, Director of the Department of Structural Biology and Chemistry. "What's more, these data need to be analyzed and processed to produce 3D images. A few days of observation by the Titan can require months of computer analysis. We need to boost the Institut Pasteur's computing power." Alongside the expert team in ultra-high resolution microscopy that will be in charge of using the microscope,* a group of scientists have been specially recruited to deal with data processing.
*The Structural Studies of Macromolecular Machines In Cellula Unit, directed by Dorit Hanein
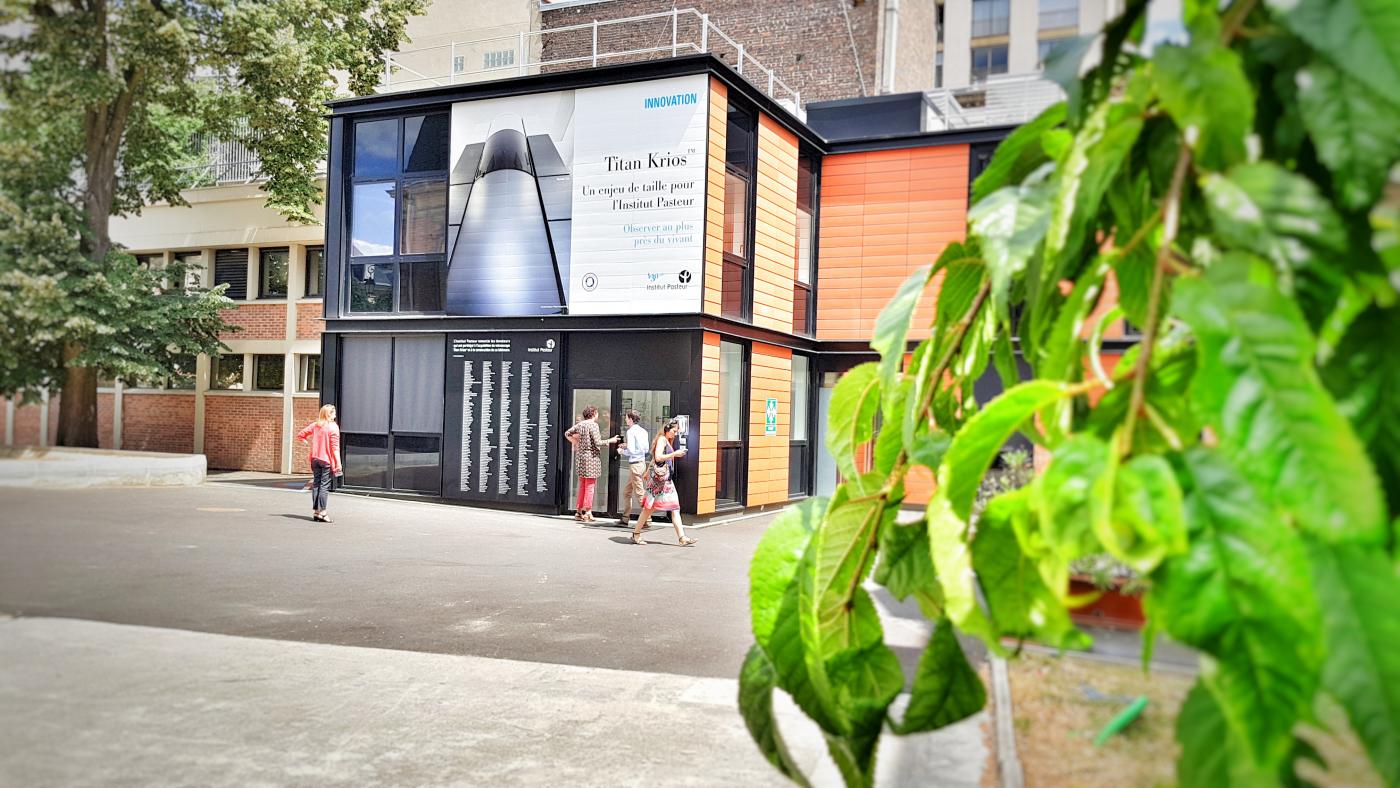

Big data is encouraging scientists to share their information
From genomic data to imaging, the quantity and quality of data collected from patients are constantly on the rise; hundreds of snippets of information are now gathered from each individual patient, compared with just a dozen a few years ago. In the United Kingdom, the UK Biobank project is collecting vast reams of health data on no fewer than 500,000 British volunteers, with the aim of "improving the prevention, diagnosis and treatment of a wide range of serious and life-threatening illnesses" – including cancer, heart diseases, diabetes, arthritis, osteoporosis, depression and dementia. This database is public and freely accessible to researchers online if they register. Users are merely asked to inform UK Biobank of discoveries made using the data "so that other scientists can benefit." This illustrates a phenomenon that is developing alongside the emergence of big data: data sharing. Researchers worldwide are now pooling their resources, working together to combine and analyze their data and providing access to the databases that they coordinate (see inset below). This has become a sine qua non to enable them to make the best use of the plethora of available information.
The world's most comprehensive database stored at the Institut Pasteur
The Institut Pasteur is to host the world's largest database on autism – not in terms of the number of patients, which currently stands at 800 and will eventually rise to 3,000, but in terms of the quantity and quality of the information gathered for each individual: brain imaging, electroencephalograms, genomic data, psychological profiles, intelligence quotient, quality of life, behavioral tests, etc. "Compared with the existing databases for autism, this takes things to a whole new level," says Thomas Bourgeron, Head of the Human Genetics and Cognitive Functions Unit at the Institut Pasteur and a pioneer in the discovery and investigation of the genes associated with autism. The scientist is coordinating the genetics part of the European program EU-AIMS* for research into new treatments for autism, which involves 106 sites in 37 countries across Europe, making it possible to collect large volumes of data. Guillaume Dumas, a member of his team, explains: "The development of this database is a technical challenge that also has to comply with the very strict ethical regulations that govern access to and security of human health data. We are working with the Information Systems Department for the storage and organization of these data, and with biostatisticians in the Center of Bioinformatics to develop tools to analyze them and facilitate access to the information for scientists." This brainwave specialist remembers how, not so long ago, he used to work "with big binders at the hospital full of hundreds of DVDs containing patients' electroencephalogram data!" The centralization of all these digitized data in the new EU-AIMS database, available online to scientists worldwide, will enable researchers to compare a huge range of information at their computers and to launch robust, reproducible analyses that will undoubtedly move our understanding of autism up several gears. And also – this is the aim – to develop treatments that are tailored to the various types of autism and the characteristics of each patient. The potential impact is huge: autism spectrum disorder affects more than 1% of the population.
*European Autism Interventions – A Multicenter Study for Developing New Medications (EU-AIMS)
Profession: Data manager
After a Master's in Bioinformatics, a PhD in Population Genomics focusing on a small cave-dwelling fish from Mexico, then a post-doctoral internship investigating another fish – this time from Cuba –, Julien Fumey decided that he wanted to take his career in a new direction. He was particularly interested in autism, and with his skills in IT and genome analysis he secured a position as data/project manager – a new profession that has emerged in the big data era –, which was advertised in January 2018 in Thomas Bourgeron's laboratory at the Institut Pasteur. "My role is to coordinate the management of data from the lab and from the EU-AIMS project (see inset above) – in terms of both storage and judicial and legal procedures," explains Julien, who acts as a new intermediary between lab scientists, hospital clinicians responsible for patient data, computer scientists and biostatisticians on campus, and especially the Legal Affairs Department and the Ethics Unit. "With the fish it was simple: I just needed to secure permission to collect and transport them," comments Julien. "When it comes to humans, I have discovered that the regulations are extremely strict, with multiple requests for permission from ethics committees, the CNIL [French Data Protection Authority], etc. All the aspects of protecting and securing patient data take a huge amount of time, especially since the Institut Pasteur is particularly rigorous in this field." The data manager's task is certainly not an easy one...

When it comes to humans, I have discovered that the regulations are extremely strict, with multiple requests for permission from ethics committees, the CNIL*, etc.
Julien Fumey Data manager in the Human Genetics and Cognitive Functions, at the lnstitut Pasteur.
* The French Data Protection Authority

Graphic interpretation of gene mapping. Credit: Adobe Stock
A sizable challenge: data storage
Another considerable challenge raised by big data – in both technical and economic terms – is how to store all this information. Research organizations are having to acquire more and more storage servers and supercomputers. The data from a single human genome can take up as much space as 100,000 holiday photos! When creating the vast European database on autism, for example (see inset above), €800,000 had to be spent just on storing patients' genomic data.
But big data holds the promise of inestimable progress in biomedical research: studies on large cohorts of patients will allow much more robust findings to be made than in the past, and access to a wealth of information about individuals gives a better idea of their specific characteristics and how their genome, microbiota, environment and lifestyle affect their response to disease, treatment and vaccines (see inset below). The ultimate aim is precision medicine: an approach that is preventive, personalized, predictive and participatory.
1,000 "healthy" volunteers for precision medicine
"What genetic, environmental and lifestyle factors influence our immune response and make us more or less likely to develop a disease or to react effectively to a given treatment or vaccine? This is the ambitious question – the key to a more personalized approach to medicine – that we are trying to answer," explains Lluis Quintana-Murci, Head of the Human Evolutionary Genetics Unit at the Institut Pasteur and coordinator of the "Milieu Intérieur" Consortium, composed of 30 teams from a range of scientific disciplines (immunology, genomics, molecular biology and bioinformatics), from a number of different research institutes and hospitals. "We are studying a cohort* of a thousand healthy volunteers aged 20 to 70, 500 women and 500 men. For each volunteer we have already gathered a considerable volume of data, and at this very moment their whole genomes are being sequenced," says the scientist. Samples of blood, saliva and stools (to investigate the gut microbiota) have been collected for each volunteer and a questionnaire with around 200 variables has been filled out: age, weight, height, medical history, sleeping habits, nutrition, smoker/non-smoker, socio-demographic data (e.g. income and level of education), etc. For each individual, some of the blood taken was divided into around thirty tubes, each of which was immediately stimulated with a pathogen (the influenza virus, the BCG, Staphylococcus, etc.) to mimic an infection and measure the immune response.

We seek to know what genetic, environmental or lifestyle factors affect our immune response and make us more or less prone to develop an illness.
Lluis Quintana-Murci Responsable de l’unité de Génétique évolutive humaine, à l’Institut Pasteur
"This broad range of data allows us to make a highly detailed analysis, with the help of computer tools, that combines environmental and socio-demographic factors with biological factors, especially the genome, so that we can obtain a very precise picture of what is involved in the variability of the immune system," adds Lluis Quintana-Murci. "The hope with big data is that we will be able to apply the knowledge derived from this study to the development of precision medicine, which involves finding the appropriate therapeutic strategy for a patient in line with his or her personal characteristics and immune profile, whether in response to infectious or inflammatory diseases or in cancer immunotherapy." Early results have already demonstrated the major impact of smoking on the composition of white blood cells, explaining why smokers are more susceptible to infection, and have enabled the scientists to identify hundreds of genetic variations that change the production of key molecules in the immune response – some of which are associated with a greater risk of developing conditions such as pollen allergy, lupus erythematosus and type 1 diabetes.
The Milieu Intérieur Consortium is a "Laboratory of Excellence" or LabEx under the French government's Investing in the Future program.
*Cohort managed by a biomedical research center in Rennes
The 10th HIV gene
Olivier Gascuel made the headlines in 2015 when, drawing on a genomic analysis of 25,000 strains of HIV,* he revealed a "hidden" gene whose existence had been suspected since the late 1980s but had been the source of much controversy. He thereby confirmed that the viral genome was indeed composed of 10 genes rather than 9, opening up new therapeutic prospects.
This pioneering bioinformatics researcher currently leads the Evolutionary Bioinformatics Unit and the Center of Bioinformatics, Biostatistics and Integrative Biology (C3BI) at the Institut Pasteur. The C3BI has a team of around 50 bioinformaticians who have been recruited since 2014 to support the teams on campus by developing IT and statistical methods to analyze their big data. "Until the late 1990s, biology was a discipline that was carried out using test tubes and Petri dishes. Nowadays, biologists are just as likely to be sat at a computer as working in a lab," notes Olivier Gascuel.
How our genes influence empathy
"Big data has called into question some things we accepted as given," explains Thomas Bourgeron, Head of the Human Genetics and Cognitive Functions Unit at the Institut Pasteur. "For example, it was widely held that individuals with autism had anomalies in a region of the brain known as the corpus callosum. The theory was put forward on the basis of studies involving small numbers of patients. However, by reproducing these studies not on 20 people but on 500 or more, this conclusion was proved to be erroneous. Nowadays, thanks to big data, our analyses are much more robust."
The scientist has been involved in a study* of 46,000 people on empathy, the ability to offer an appropriate emotional response to the thoughts and feelings of others. The findings suggest that our empathy is not just the result of our upbringing and experience, but is also partly influenced by our genes: a tenth of the variation in the degree of empathy between individuals is thought to be down to genetic factors...
*led by scientists from the University of Cambridge, the Institut Pasteur, Paris Diderot University, the CNRS and the genetics company 23andMe
The Neanderthal legacy and immunity
"The emergence of big data has opened doors for us in a remarkable way," observes Lluis Quintana-Murci, Head of the Human Evolutionary Genetics Unit. "For example, nowadays we can study the innate immune system, our first line of defense against pathogens, as a whole. Whereas not long ago we were analyzing two or three genes involved in this immunity, now we are studying more than 1,500." His team recently demonstrated that the genes involved in our innate immune system were enriched by the legacy of Neanderthals. This knowledge of human evolutionary history – much like the comparison of the current genomes of different populations (Europeans, Africans, Asians, etc.) – can help us to identify the most important genes, those that have enabled us to adapt the best and survive pathogens. "Big data has revolutionized our research on genome biodiversity. Studying the genetic variability of populations is vital in helping us understand our inequalities when it comes to disease and also our response to treatments and vaccines."
Joining forces to shed light on disease emergence. These three scientists and several other teams at the Institut Pasteur are involved in the large-scale INCEPTION project, launched in 2016 under the French government's Investing in the Future program. INCEPTION combines integrative biology, social science and data analysis techniques to study the emergence of infectious diseases (Ebola, Zika, HIV/AIDS, etc.) and other conditions (cancer, autism, etc.) in individuals and populations.