Scientists at the Institute Pasteur and CNRS have identified key regulatory factors controlling one of the critical developmental processes occurring during embryo development : X-inactivation, which ensures the silencing of the genes carried by one of the two X chromosomes present in all cells of female mammals. These regulatory factors are also implicated in maintaining the capacity of embryonic stem cells to give rise to the different tissues which form our organism, such as the skin, liver and brain. And these same regulatory factors are also capable of ‘reprogramming' the genomes of adult cells so that they lose their specialisation and return to the stem cell state. The research published in Nature contributes to our understanding of the fundamentals governing the stem cell state, a knowledge which will be necessary for controlling the differentiation of these cells and developing their use as a base for novel therapeutic stratégies.
Press release
Paris, november 17, 2010
Female mammals carry two identical sex chromosomes, two X chromosomes, whilst males possess only a single X chromosome. To avoid the effects of this imbalance, a mechanism is activated in females during embryonic development which ensures the silencing of the genes present on one of the two X chromosomes present in each cell. This mechanism ensures that cells of males and females express equally the genes present on the X chromosome.
The mechanisms responsible for this X-chromosome inactivation are under investigation in the unit of Mouse Molecular Genetics (Institut Pasteur/CNRS URA 2578), headed by Philip Avner. The scientists in this unit have previously identified three factors controlling directly the onset of this process in the embryo. In collaboration with the team of Dr.Ian Chambers at the University of Edinburgh, they have now characterised a further three factors implicated in a complementary second level of regulation.
Interestingly some of these factors have previously been shown to be able to induce adult cells of specialised tissues such as the skin to move towards the undifferentiated or stem cell state. When this ‘deprogrammation’ occurs in female cells it is accompanied by the reactivation of genes on the inactive X chromosome. This suggests that there are common molecular mechanisms between the reactivation of the inactive X and the process of reprogrammation/deprogrammation.
The discovery of scientists from the Institut Pasteur and CNRS by contributing to the identification of these underlying common regulatory mechanisms. underlines the importance of fundamental research for our understanding of the nature of the developmental plasticity of embryonic stem cells, an area of potential major interest for the development of novel therapeutics and public health.
_ _ _
These works received financial supports from the Agence Nationale de la Recherche Scientifique and the Epigenome European Network of Excellence.
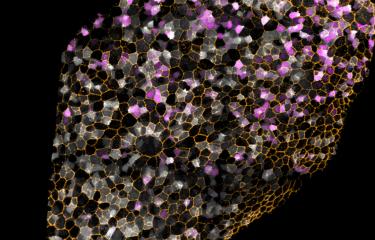
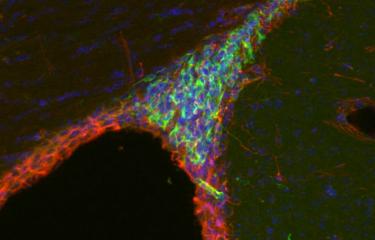
Picture : The inactivated X-chromosom appears in green, via the non-coding ARN Xist, which covers it in the nucleus of each cell of female mammals. © Institut Pasteur
Source
Molecular coupling of Tsix regulation and pluripotency, Nature, 17 novembre 2010.
Pablo Navarro (1,2), Andrew Oldfield (1,3), Julie Legoupi (1), Nicola Festuccia (2), Agnès Dubois (1), Mikael Attia (1), Jon Schoorlemmer (4), Claire Rougeulle (1,3), Ian Chambers (2) and Philip Avner (1).
(1) Unité de Génétique Moléculaire Murine, URA 2578, Institut Pasteur 75724 Paris Cedex 15, France.
(2) Medical Research Council (MRC) Centre Development in Stem Cell Biology, Institute for Stem Cell Research, School of Biological Sciences, University of Edinburgh, Edinburgh EH9 3JQ, UK.
(3) UMR 7216 Epigénétique et Destin Cellulaire, CNRS/Université Paris Diderot, Case 7042, 75205 Paris Cedex 13, France.
(4) ARAID Foundation and Instituto Aragonés de Ciencias de la Salud, DepartamentoAnatomía y Embriología, Facultad de Veterinaria, 50013 Zaragoza, Spain.
Contact
Institut Pasteur Press Office
Marion Doucet - + 33 (0)1 45 68 89 28 - marion.doucet@pasteur.fr
Nadine Peyrolo - + 33 (0)1 45 68 81 47 - nadine.peyrolo@pasteur.fr