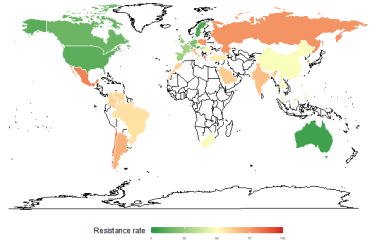
Designing effective new molecules to tackle infectious diseases is a difficult process that currently represents a huge challenge for scientists, especially given the rise in bacterial resistance to antibiotics. Scientists from the Institut Pasteur's Bacterial Genome Plasticity Unit have responded to this challenge by developing a new biotechnological tool which bypasses the sampling limitations that represent a hurdle for scientists, paving the way for the identification and use of effective new molecules.
There are two types of molecular machinery that produce antibiotics: polyketide synthases (PKSs) and non-ribosomal peptide synthetases (NRPSs). Since the 1940s, the pharmaceutical industry has successfully searched multiple PKS and NRPS libraries with a fine-tooth comb. But this system has now reached its limits – all the therapeutic molecules have been found.
Overcoming the limits of recombination
So the Institut Pasteur's scientists have turned to another approach: recombination, a naturally occurring mechanism in all living organisms in which genes are mixed into new combinations. In biotechnology, where recombination is triggered artificially (one DNA molecule is cut and joined to another), it is an effective tool to generate new genetic combinations and promote the diversity of the molecules produced. But it has its limits:
- in "sequence-specific" recombination systems, the recombined sequence cannot easily be changed or has to stay constant;
- in "homologous" recombination systems, the target sites (DNA regions) are highly similar.
These structural limitations reduce the possibilities of inserting recombined DNA into DNA regions that already have a function. "Developing a recombination system that is "site-specific" rather than "sequence-specific" means that recombination sites can be integrated into practically any DNA sequence," explains Didier Mazel, Head of the Bacterial Genome Plasticity Unit at the Institut Pasteur.
A biotechnology tool based on the integron system
The members of Didier Mazel's team have focused their efforts on the integron, a genetic element known for its role in a recombination mechanism. "By observing integron recombination sites, we realized that there was very low specificity in the sequences – just 3 or 4 bases [small molecules that compose DNA] –, and also that the distance between them was variable," explains Didier Mazel.
The scientists determined the recognition characteristics of the recombination system: it occurs on single-stranded DNA, and the secondary structure of the DNA rather than the primary structure is recognized (in other words a few bases outside the DNA helix structure). By elucidating the mechanism of operation, the recombination potential of integrons can be harnessed: "We can artificially create gene combinations that do not exist in nature and may enable the effective synthesis of new molecules," continues Didier Mazel. There is currently a pressing need to produce new molecules to tackle the antibiotic resistance crisis.
Using machine learning in biotechnology
In connection with this research, the scientists had to design a computer program that enabled the de novo creation of genetic sequences with recombination potential. "We started out thinking that we knew all there was to know about the recombination determinants in these genetic elements that we have been working on for more than 20 years. But when we tested our genetic sequences, we realized that some recombined well, others fairly well, and still others not at all," explains Didier Mazel. Working with Julia Bernardes from the Laboratory of Computational and Quantitative Biology at the Sorbonne University Association, his team attempted to understand what influenced these variations by applying a machine learning approach to extract information by testing different theories. "We discovered new determinants that improve our ability to predict the recombination efficacy of de novo sequences at the sites," concludes the scientist.
Source
Structure-specific DNA recombination sites: design, validation and machine learning based refinement, Science Advances, July 24th, 2020
Aleksandra Nivina1,2,3, Maj Svea Grieb1,2, Céline Loot1,2, David Bikard1,2, Jean Cury1,2,3, Laila Shehata1,2, Juliana Bernardes4 and Didier Mazel1,2
1 Unité Plasticité du Génome Bactérien, Institut Pasteur, 75724 Paris, France
2 CNRS UMR3525, 75724 Paris, France
3 Paris Descartes, Sorbonne Paris Cité, Paris, France
4 Laboratoire de Biologie Computationnelle et Quantitative, Sorbonne Universités, CNRS 19 UMR7238, 75005 Paris, France.
This study is part of the priority scientific area Antimicrobial Resistance of the Institut Pasteur's strategic plan for 2019-2023.