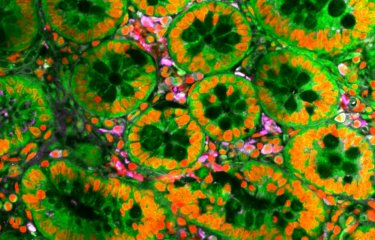
Candida albicans is one of the most formidable fungal species, causing infection in humans. Investigating the structure and reproduction methods of pathogenic populations can help us to understand how they emerge and spread. A team of scientists therefore decided to sequence and analyze the genomes of 182 strains of C. albicans from around the world. They confirmed the clonal reproduction of this human pathogen but also showed that parasexual reproduction, previously only observed in a laboratory setting, contributes to the genetic diversity of C. albicans and therefore also to its ability to adapt to new environments and rid itself of deleterious mutations.
There are 5 million fungal species, but only a few hundred can cause disease in humans. Candida albicans is one of the most formidable of these species. It belongs to one of the four genera of pathogenic fungi responsible for high mortality rates in humans and is the second most common agent of opportunistic fungal infection in the world. Candida albicans is part of the human gut microbiota (a commensal fungus) but it also causes mucosal infections in healthy individuals and severe opportunistic infections in those with weakened immune defenses (immunocompromised individuals and patients who have received organ transplants, undergone serious surgery or suffered major trauma).
Understanding how pathogens emerge and spread involves analyzing the structure of their populations. Several studies have shown the importance of population genetics in shedding light on the emergence of new diseases, such as white-nose syndrome in bats in North America, which is caused by a fungus and is devastating entire bat populations. In this case, population genetics studies revealed that a fungus of European origin, which is harmless in European bat populations, invaded North America via clonal expansion.
Scientists in the Biology and Fungal Pathogenicity Unit at the Institut Pasteur and INRA, in collaboration with 12 other teams, sequenced and analyzed the genomes of 182 strains of C. albicans isolates, either commensal or responsible for superficial or invasive infections, from around the world. This is the largest population genomics study carried out on the pathogen to date. It confirms the primarily clonal reproduction of this human pathogen. But it also shows traces of introgression in the genome of some strains, indicating the possibility of genetic exchanges between strains in nature and reflecting parasexual reproduction, independent from meiosis, which had previously only been observed in a laboratory setting, or sexual reproduction, hitherto unknown for C. albicans. C. albicans' use of (para)sexual reproduction is no doubt crucial for it to generate genetic diversity and adapt to new environments quickly, as well as to rid itself of the deleterious mutations that build up during clonal reproduction and which, if they are not eliminated, would lead to the species' extinction.
Candidiasis
Fungi (yeast) of the Candida genus can cause superficial infections affecting the mucous membranes or skin, and invasive infections, either localized to one organ or generalized throughout the body. Of the 200 known species of Candida, around twenty are responsible for human infection. Candida yeasts are often responsible for severe, hospital-acquired infections.
Read the fact sheet about "Candidiasis" (page in French)
Source
Gene flow contributes to diversification of the major fungal pathogen Candida albicans, Nature Communications, June 8, 2018.
Jeanne Ropars1,2, Corinne Maufrais1,3, Dorothée Diogo1, Marina Marcet-Houben1,4,5, Aurélie Perin1, Natacha Sertour1, Kevin Mosca1, Emmanuelle Permal1, Guillaume Laval3,6, Christiane Bouchier7, Laurence Ma7, Katja Schwartz8, Kerstin Voelz9, Robin C. May9, Julie Poulain10,11,12, Christophe Battail10, Patrick Wincker10,11,12, Andrew M. Borman13, Anuradha Chowdhary14, Shangrong Fan15, Soo Hyun Kim16, Patrice Le Pape17, Orazio Romeo18,19, Jong Hee Shin16, Toni Gabaldon4,5,20, Gavin Sherlock8, Marie-Elisabeth Bougnoux1,21,22 & Christophe d’Enfert1
1. Department of Mycology, Fungal Biology and Pathogenicity Unit, Institut Pasteur, INRA, 75015 Paris, France.
2. Ecologie Systématique et Evolution, CNRS, Univ. Paris Sud, AgroParisTech, Université Paris Saclay, 91405 Orsay cedex, France.
3. Center for Bioinformatics, BioStatistics and Integrative Biology (C3BI), USR 3756 IP CNRS, Institut Pasteur, 75015 Paris, France.
4. Centre for Genomic Regulation (CRG), The Barcelona Institute for Science and Technology, 08003 Barcelona, Spain.
5. Universitat Pompeu Fabra (UPF), 08002 Barcelona, Spain.
6. Department of Genomes and Genetics, Human Evolutionary Genetics Unit, UMR 2000 CNRS, Institut Pasteur, 75015 Paris, France.
7. Biomics Pole, CITECH, Institut Pasteur, 75015 Paris, France.
8. Department of Genetics, Stanford University Medical School, Stanford, CA 94305-5120, USA.
9. School of Biosciences and Institute of Microbiology and Infection, University of Birmingham, Birmingham B15 2TT, UK.
10. CEA, Genoscope, Institut de biologie François Jacob, 91000 Evry, France.
11. CNRS UMR 8030, 91000 Evry, France.
12. Univ. Evry, Univ. Paris-Saclay, 91000 Evry, France.
13. UK National Mycology Reference Laboratory, Public Health England, Bristol BS2 8EL, UK.
14. Department of Medical Mycology, Vallabhbhai Patel Chest Institute, University of Delhi, Dehli 110007, India.
15. Department of Obstetrics and Gynecology, Peking University Shenzhen Hospital, PR Guangdong Sheng 518036, China.
16. Department of Laboratory Medicine, Chonnam National University Medical School, Gwangju 61469, South Korea.
17. EA1155 – IICiMed, Institut de Recherche en Santé 2, Université de Nantes, 44200 Nantes, France.
18. Department of Chemical, Biological, Pharmaceutical and Environmental Sciences, University of Messina, 98166 Messina, ME, Italy.
19. IRCCS – Centro Neurolesi Bonino-Pulejo, 98124 Messina, Italy.
20. ICREA, 08010 Barcelona, Spain.
21. Unité de Parasitologie-Mycologie, Service de Microbiologie clinique, Hôpital Necker-Enfants-Malades, Assistance Publique des Hôpitaux de Paris (APHP), 75015 Paris, France.
22. Université Paris Descartes, Sorbonne Paris-Cité, 75006 Paris, France.