The Institut Pasteur is at the forefront of computational biology research. Scientists in the Evolutionary Bioinformatics Unit (part of Institut Pasteur and CNRS Unit, USR 3756) recently developed the PastML program. This tool, based on decision-theory concepts, infers "ancestral scenarios" that describe the evolution of traits or characters along phylogenetic trees. The characters can be very diverse and represent the morphology of species studied, biochemical properties of proteins that evolved from a common ancestral protein, the geographical origin of an outbreak and its global spread, or the emergence and spread of resistance to treatments. In mere minutes, PastML analyzes vast reams of data and suggests the most likely hypotheses in an accurate and robust manner. PastML is available to the international scientific community, just as Phylogeny.fr since 2008 and now NGPhylogeny.fr, a new version entirely rebuilt by the same teams in 2019.
The reconstruction of ancestral scenarios is widely used by scientists to study the evolution of characters along phylogenetic trees. "This is the case, for example, with morphological, geographical, ecological or molecular characters, for which large amounts of data exist, due largely to sequencing techniques", explains Olivier Gascuel, Director of the Department of Computational Biology at the Institut Pasteur. Pathogens are a preferred application of these reconstructions, which enable us to travel back in time and study the ancestral sequences that caused pandemics, along with their geographical origins and the different hosts and their transmission to humans.
Until recently the scientists used this data with Bayesian methods which require extremely long computation times and are ill-suited to large samples, or else used maximum likelihood (ML) methods, which are faster but incapable of reflecting data uncertainty simply and concisely. ML methods either produced probability vectors that were difficult to interpret (marginal reconstructions), or single decisions (joint reconstructions) with no indication of alternatives. Biologists’ intuition is insufficient for understanding and interpreting these complex results, and mathematics and statistics are needed to support, confirm, reorient or invalidate the hypotheses made.
This was the idea behind the simple and fast approach proposed by the Institut Pasteur’s computational biology experts which draws on decision-theory concepts (notably the Brier score invented for weather forecasts): "We associate each node in the phylogenetic tree with a set of likely character states; just one in the simple tree nodes, generally near the leaf nodes and recent information, and several in the difficult regions, typically the oldest ones near the root node", sums up Gascuel. "The method combines three important aspects: decision theory, computation speed, and visualization via graphic tools and user-friendly interfaces."
Dengue and HIV, conclusive initial results
In collaboration with the University of Tokyo, the method was implemented in a computer program called PastML, which the team tested on several datasets and epidemiological issues. PastML was applied to the evolution of drug resistance in a large HIV dataset, and to the phylogeography of dengue serotype 2 (DENV2). "These analyses took mere minutes and produced convincing results, demonstrating the approach’s accuracy and robustness."
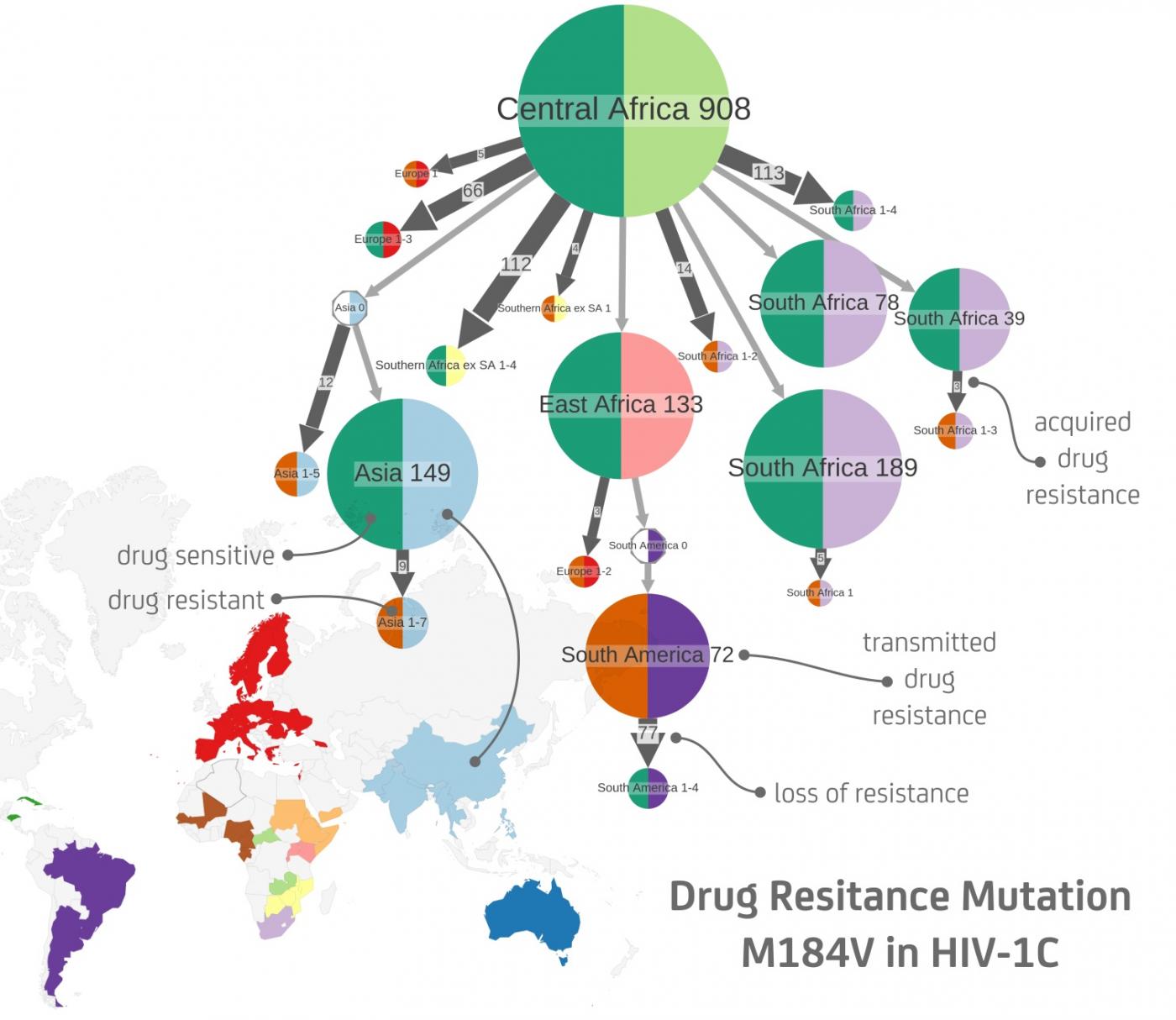
- For HIV, the scientists reconstructed the evolution of treatment resistance in the population since the arrival of the first drugs in the mid-1990s. It is generally considered that resistance mutations emerge independently under treatment pressure, and subsequently that resistance clusters appear, corresponding to transmission between untreated patients. These resistance clusters are particularly problematic, raising fears of emergence of multi-resistant strains that are resistant to all drugs, as is the case with other pathogens (notably malaria). "What we see is consistent with the theory, explains Olivier Gascuel. Based on these results, instead of the previous qualitative approach we now have a quantitative one to visualize, predict and monitor resistant sub-epidemics observed in particular regions or within groups of patients presenting specific risks."
- For dengue (DENV2), the scientists reconstructed the phylogeography scenario explaining the emergence of this pandemic in humans and its global spread. On the whole, these results were in keeping with previous knowledge – its endemic and epidemic origins can be traced back to South East Asia around 1750, and its spread across the continents and sub-continents is consistent with other analyses, but the results also show that the ancestral root of sylvatic strains, which predates the human pandemic, cannot be elucidated with the data available today.
The Institut Pasteur in support of open science
The PastML program proved its efficacy and is now available to the international community, once again demonstrating the Institut Pasteur's expertise in bioinformatics and its dedication to producing open software. The Paris campus is leading the way in computational biology research, most notably with its new dedicated scientific department set up in spring 2019, developed from a center originally established in 2015. The department represents a new phase for the Institut Pasteur and reflects the leading role it plays in this research field.
Source :
A Fast Likelihood Method to Reconstruct and Visualize Ancestral Scenarios, Mol. Biol. Evol., May 24, 2019
Sohta A. Ishikawa *,1,2,3, Anna Zhukova *,1, Wataru Iwasaki 2, and Olivier Gascuel1
1. Unité Bioinformatique évolutive, Institut Pasteur, C3BI USR 3756 IP & CNRS, Paris, France
2. Department of Biological Sciences, The University of Tokyo, Tokyo, Japan
3. Evolutionary Genomics of RNA Viruses, Virology Department, Institut Pasteur, Paris, France
* These authors contributed equally to this work.
NGPhylogeny.fr: the 2019 version of a widely used open science program
Phylogeny.fr, developed in 2008, was designed to facilitate the execution of phylogenetic workflows, and is nowadays widely used. "The international scientific community makes extensive use of it for highly diverse analyses and issues. It was cited more than 3,000 times in the space of 10 years", explains Olivier Gascuel. However, since its development user needs have changed, new tools and workflows have been published, and the number of jobs submitted has risen dramatically, thus promoting new practices, which motivated its refactoring.
In April 2019, the teams in the Institut Pasteur's Department of Computational Biology released a completely revised version. They developed NGPhylogeny.fr to allow for greater flexibility in terms of both tools* and their workflows via a dynamic and user-friendly interface. These tools have a wide variety of uses (sequence searching, multiple sequence alignment, model selection, phylogenetic tree inference and tree viewing), and cover a large number of methods (distance, parsimony, maximum likelihood and Bayesian methods). They are supported in preconfigured workflows with multiple options:
- "One Click", for non-experts;
- "Advanced", i.e. customizable for certain criteria;
- "A la Carte", built from scratch, for specialists only.
NGPhylogeny.fr is a scalable solution that is easily deployable on any server or personal computer. "The Bioinformatics and Biostatistics Hub – the Institut Pasteur’s campus-based service platform – played a major role in NGPhylogeny.fr's development. This web server is of great service to the international community; it is already used daily by more than 100 researchers worldwide. It's a fantastic showcase for our services and expertise."
*NGPhylogeny.fr features the latest versions of a number of tools (TNT, FastME, MrBayes, etc.) as well as new tools designed over the past decade (PhyML 3.0, SMS, FastTree, trimAl, BOOSTER, etc.).
Source :
NGPhylogeny.fr: new generation phylogenetic services for non-specialists, Nucleic Acids Research, July 2, 2019
Frédéric Lemoine 1,2,*, Damien Correia 1,3,4, Vincent Lefort 3, Olivia Doppelt-Azeroual 2, Fabien Mareuil 2, Sarah Cohen-Boulakia 4,* and Olivier Gascuel 1,3,*
1. Unité Bioinformatique évolutive, C3BI USR 3756, Institut Pasteur & CNRS, Paris, France,
2. Hub Bioinformatique et Biostatistique, C3BI USR 3756, Institut Pasteur & CNRS, Paris, France,
3. Méthodes et Algorithmes pour la Bioinformatique, LIRMM UMR 5506, Université de Montpellier & CNRS, Montpellier, France
4. Laboratoire de Recherche en Informatique, Université Paris-Sud, CNRS UMR 8623, Université Paris-Saclay, Orsay, France