Phage therapy, which uses viruses known as bacteriophages to treat bacterial infections, is a long-standing medical procedure whose mechanisms of action are still poorly understood. Scientists from the Institut Pasteur and CNRS have demonstrated in vivo in a murine model that bacteria are capable of regulating their gene expression to evade the numerous bacteriophages present in the gut environment. This research explains the difference in bacteriophage efficacy between in vitro and in vivo conditions. The findings were published in the journal Cell Host & Microbe on April 13, 2022.
Phage therapy is a medical approach that involves treating bacterial infectious diseases using the natural ability of certain viruses, known as bacteriophages, to kill bacteria that they specifically recognize. A significant decline in the use of this therapeutic strategy discovered over 100 years ago was seen in the West following the development of antibiotics. However, faced with an alarming increase in the number of infections caused by antibiotic-resistant bacteria and the concerning prospect of being left with no treatment options, scientists are seeking to shed light on bacteriophages' mechanism of action.
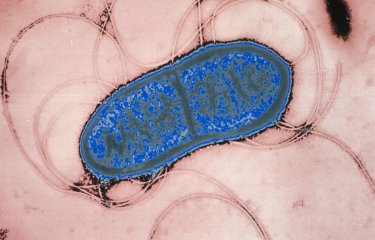
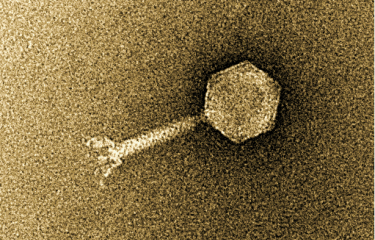
Bacteria and bacteriophages are the most abundant entities in the human gut microbiota. Although bacteriophages kill bacteria, the two antagonist populations coexist in a balance in the gut.
Until now, insufficient scientific data have been available to understand how phage therapy works in vivo. Interactions between bacteria and bacteriophages have, in contrast, been extensively studied in vitro. In these conditions, bacteriophages quickly infect bacteria, replicate, and destroy bacteria, while releasing new viruses capable of infecting other bacteria. However, the dynamics observed between these two microorganisms are very different in the gut of mammals . Some bacteriophages that are effective in culture medium are totally ineffective in the gut environment.
In order to understand this difference, scientists at the Institut Pasteur and CNRS decided to compare the gene expression profile, or transcriptome, of the bacterium Escherichia coli in both contexts: culture media and the gut. Using this method, they revealed genetic regulations that characterize the bacterium's adaptation to the gut environment.
By closely examining the genes involved in this adaptation, they revealed four genes that modulate the bacterium's susceptibility to bacteriophages. "We observed that certain genes required for infection by bacteriophages are expressed less in the gut than in vitro, thus protecting bacteria from bacteriophages," commented Laurent Debarbieux, Head of the Bacteriophage, Bacterium, Host Unit at the Institut Pasteur (CNRS joint unit)[1] and last author of the study. The scientists were able to verify their theory by eliminating the expression of one particular gene. They observed that bacterial susceptibility to a bacteriophage was significantly reduced. Consequently, bacteria in the gut are able to resist predation by bacteriophages by modulating the expression of certain genes rather than mutating their genome.
This study therefore demonstrates that environment plays a predominant role in interactions between bacteria and bacteriophages. These findings pave the way for improved use of bacteriophages for therapeutic purposes.
[1] The CNRS research unit's name is the "Integrative and Molecular Microbiology Unit".
Source
The gut environment regulates bacterial gene expression which modulates susceptibility to bacteriophage infection, Cell Host & Microbe, 13 avril 2022
Marta Lourenço1,2#, Lorenzo Chaffringeon1,3,4, Quentin Lamy-Besnier1, Marie Titécat1,5, Thierry Pédron1, Odile Sismeiro6, Rachel Legendre6,7, Hugo Varet6,7, Jean-Yves Coppée6, Marion Bérard8, Luisa De Sordi1,3,4 and Laurent Debarbieux1*
1 Institut Pasteur, Université Paris Cité, CNRS UMR6047, Bacteriophage Bacterium Host, Paris F-75015 France
2 Sorbonne Université, Collège Doctoral, F-75005 Paris, France
3 Sorbonne Université, INSERM, Centre de Recherche St Antoine, UMRS_938, Paris, France
4 Paris Center for Microbiome Medicine (PaCeMM) FHU, AP-HP, Paris, Ile-de-France, France
5 Université de Lille, INSERM, CHU Lille, U1286-INFINITE-Institute for Translational Research in Inflammation, F-59000 Lille, France
6 Transcriptome and EpiGenome Platform, Biomics, Center for Technological Resources and Research (C2RT), Institut Pasteur, Université Paris Cité, Paris F-75015 France
7 Bioinformatics and Biostatistics Hub, Department of Computational Biology, Institut Pasteur, Université Paris Cité, Paris F-75015 France
8 Institut Pasteur, Université Paris Cité, DT, Animalerie Centrale, Centre de Gnotobiologie, 75724 Paris, France
# present address: Department of Global Health, Institut Pasteur, Université Paris Cité, Paris F-75015 France
* Lead author and correspondence