A study carried out by Institut Pasteur researchers in cooperation with Inserm and INRA reveals how Listeria monocytogenes, the bacterium responsible for listeriosis, changes the activity of its entire genome to shift from an inoffensive to a pathogenic state. New types of gene products (RNAs) have been discovered. This research, which appeared in the advanced online edition of the journal Nature, will be published in the journal on June 18, 2009.
Press release
Paris, june 11, 2009
Institut Pasteur researchers from the Bacteria-Cell Interactions Unit (Inserm Unit 604, INRA USC2020), led by Pascale Cossart, in cooperation with other groups from the Institut Pasteur (Microbes and Host Barriers Group, Inserm Avenir 604; In Silico Genetics Unit, Institut Pasteur Genopole) and a Swedish team, have demonstrated how this inoffensive bacterium in the environment becomes a dangerous pathogen in its mammal host.
Using new DNA micro-arrays, which enable the entire genome of the bacterium to be analyzed (genes and inter-gene regions), the scientists compared the activity of the genome in its inoffensive state and its pathogenic state. To do this, they analyzed all the gene products in the bacterium (RNAs) and studied their regulation when the bacterium grows in the environment or in the infected host (in the intestine or blood). The set of all bacterial RNAs in a genome at a given time is known as the transcriptome.
The results of this comprehensive "transcriptomic" analysis firstly showed that when the bacterium reaches the intestine, and then the bloodstream, it radically changes the activity of its genome and successively activates various groups of virulence genes. 50 small RNAs were also identified. Some are absent from the non-pathogenic species Listeria innocua and are differentially expressed during infection; at least two of these small RNAs contribute to the virulence of Listeria monocytogenes. The researchers also discovered a series of new types of regulatory RNAs (for example long non coding antisense RNAs) which certainly also exist in other bacteria.
This work paves the way for a full understanding of Listeria's adaptation mechanisms when the bacterium moves from the environment to the infected host. They especially open up new possibilities in the highly competitive field of RNA regulation, in all species of the living world.
--
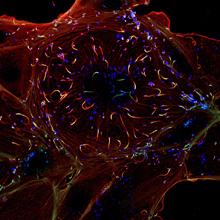
Observation by immunofluorescence of a cell (red and green) infected by Listeria monocytogenes (blue). Copyright Institut Pasteur.
Source
The Listeria transcriptional landscape from saprophytism to virulence, Nature, 18 juin 2009.
Alejandro Toledo-Arana(1,2,3), Olivier Dussurget(1,2,3), Georgios Nikitas(2,4,5), Nina Sesto(1,2,3), Hélène Guet-Revillet(1,2,3), Damien Balestrino(1,2,3), Edmund Loh(6,7), Jonas Gripenland(6,7), Teresa Tiensuu(6,7), Karolis Vaitkevicius(6,7), Mathieu Barthelemy(8), Massimo Vergassola(9,10), Marie-Anne Nahori(1,2,3), Guillaume Soubigou(8), Béatrice Régnault(8), Jean-Yves Coppée(8), Marc Lecuit(2,4,5,11), Jörgen Johansson(6,7) & Pascale Cossart(1,2,3).
(1) Institut Pasteur, Bacteria-Cell Interactions Unit
(2) Inserm, U604
(3) INRA, USC2020
(4) Institut Pasteur, G5 Microbes and Host Barriers
(5) Inserm, Avenir, U604
(6) Department of Molecular Biology
(7) The Laboratory for Molecular Infection Medicine Sweden (MIMS), Umeå University, S-90187 Umeå, Sweden
(8) Institut Pasteur, Genopole, Platform 2
(9) Institut Pasteur, In Silico Genetics Candidate Unit
(10) CNRS, URA2171
(11) Paris Descartes University, Necker Enfants Malades Hospital and Paris Public Hospital Network (AP-HP)
Read the article online on Nature website.
Press contacts
Institut Pasteur Press Office
Marion Doucet or Nadine Peyrolo – + 33 (0)1 45 68 89 28 - presse@pasteur.fr
Inserm Press Office
Anne Mignot – + 33 (0)1 44 23 60 73 - presse@inserm.fr
Inra Press Office
Mathilde Maufras – + 33 (0)1 42 75 91 69 – presse@inra.fr


