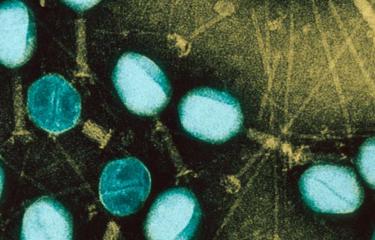
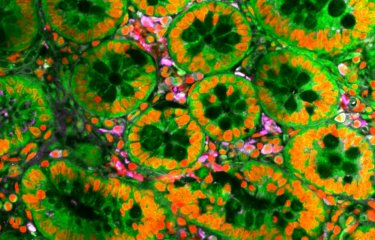
In a pioneering work, a multidisciplinary team of researchers from “Miguel C. Rubino” Lab at the Ministry of Livestock (MGAP), the National Agricultural Research Institute (INIA), the state university (Udelar) the Institut Pasteur in Montevideo and the Institut Pasteur (Paris) have identified for the first time the pathogenic strains of the Leptospira bacteria that circulate in Uruguay infecting cattle and people.
This bacteria is the cause of Leptospirosis (read the Institut Pasteur disease fact sheet), an infectious disease that has a high economic impact in the livestock sector, and it is often transmitted to humans, in particular rural and slaughterhouses workers, veterinarians, among others. The infection occurs through water that can initially be contaminated through the urine of infected animals, including rats, cattle, horses, sheep, pigs and dogs. In Uruguay, beef and dairy exports are leading sources of national income.
Because different strains of the bacteria vary in their ability to cause disease and to jump between species, it was important to identify the Leptospira variants that infect cattle in Uruguay. With this aim, a multicentric consortium was created involving the Institut Pasteur in Montevideo, the Faculty of Medicine (Udelar), INIA and the veterinary lab at MGAP.
This multidisciplinary team sampled urine and blood from 963 cattle at 48 beef and dairy farms around Uruguay. Additionally, they collected the urine and kidneys from 577 animals from 22 slaughterhouses. Each sample was tested for the presence of Leptospira and, if present, for the exact strain.
The researchers found that 20% of all cattle sampled were shedding pathogenic Leptospira in their urine, representing a large public health risk. Around 40 different strains of the bacteria were isolated, uncovering an unexpectedly large variation. The bacteria identified included three rare isolates undetected by normal tests, and two serotypes of the bacteria that matched exactly with those previously isolated from leptospirosis patients.
“This report of local Leptospira strains shall improve diagnostic tools and the understanding of leptospirosis epidemiology in South America,” Alejandro Buschiazzo, from IP Montevideo and corresponding author of the article. “These strains could also be used as new components within bacterin vaccines to protect against the pathogenic Leptospira strains that are actually circulating, a direct measure to reduce the risk of human leptospirosis.”
A project with highly trained human resources
The multicentric project was aimed at creating a bank of strains of Leptospira isolated from infected cattle in Uruguay. Each of the institutions involved has provided complementary capabilities, from obtaining samples from cattle (from farm to slaughterhouses), to the microbiological isolation and identification of strains of pathogenic bacteria.
The Biology of Spirochetes Unit (Mathieu Picardeau) at the Institut Pasteur, in Paris, works in tight collaboration with the Laboratory of Molecular & Structural Microbiology (Alejandro Buschiazzo) at the Institut Pasteur de Montevideo through a Pasteur international joint unit called Integrative Microbiology of Zoonotic Agents (IMiZA). For this work published in PLoS NTD, the Biology of Spirochetes Unit has provided expertise and training to the Montevideo staff, for the identification and typing of Leptospira strains. This has led to ongoing work analyzing whole genome sequences of animal and human strains, which will be informative in the formulation of more efficacious vaccines.
The first phase of the project consisted in the isolation and typing of the strains, in addition to the development of the molecular techniques necessary for the work. As a result, the groud created a bacterial bank that already contains more than 60 strains. Now, at the beginning of the second phase, the consortium will seek to consolidate the epidemiological information at a national level, as well as to continue with the development of more specific vaccines.
Thanks to this joint experience, highly trained human resources have been trained in the country, integrating skills in infectious diseases of large animals, microbiology, molecular biology, and immunology towards the development of vaccines.
In this work, the National Agency for Research and Innovation (ANII) has been a fundamental actor as the main funding of the project, which also had resources from INIA and companies linked to the animal health sector (Virbac Uruguay SA, Prondil and Microsules Laboratories).
Leptospirosis: why it matters
Leptospirosis, one of the most widespread zoonoses worldwide, is a transmissible disease that affects animals and humans, caused by the pathogenic species of the genus Leptospira.
In cattle, acute infection of calves causes septicemia (read the Institut Pasteur disease fact sheet - in French) and high mortality, and in cows it triggers abortions, birth of weak offsprings, mastitis and agalactia (decrease or lack of breast milk after birth). Chronic infection also reduces the reproductive capacity of the herd and increases the interval between births.
In humans, the infection can be acquired through the contaminated environment or directly from contact with the urine of infected animals, and the originating disorder can present severe clinical signs. Although leptospirosis responds well to antibiotic treatment, if it is not diagnosed and treated properly, it can be fatal.
Antibiotics should be used to cure the disease, both in animals and in humans. Although there are some vaccines available for the preventive control of leptospirosis in animals —which entail indirect protection for humans—, the antigenic variability of the genus Leptospira results in a great variety of strains, which impose an important challenge for the diagnosis , and especially for the formulation of effective vaccines, which are made with inactivated bacteria.
To provide protection, vaccines must include the variants of strains that actually circulate in a given region, since the use of different varieties does not confer proper protection.
Source
Isolation of pathogenic Leptospira strains from naturally infected cattle in Uruguay reveals high serovar diversity, and uncovers a relevant risk for human leptospirosis, PLOS Neglected Tropical Diseases, September 13, 2018.
Leticia Zarantonelli 1,2*, Alejandra Suanes 3*, Paulina Meny 4*, Florencia Buroni 5*, Cecilia Nieves 1**, Ximena Salaberry 3**, Carolina Briano 6**, Natalia Ashfield 4, Caroline Da Silva Silveira 7, Fernando Dutra 6, Cristina Easton 3, Martin Fraga 7, Federico Giannitti 7, Camila Hamond 2,7, Melissa Macias-Rioseco 7, Clara MeneÂndez 4, Alberto Mortola 3, Mathieu Picardeau 8,9, Jair Quintero 4, Cristina Rios 4, Victor Rodriguez 5, Agustin Romero 6, Gustavo Varela 4, Rodolfo Rivero 5*, Felipe Schelotto 4*, Franklin Riet-Correa 7*, Alejandro Buschiazzo 1,9,10*, on behalf of the Grupo de Trabajo Interinstitucional de Leptospirosis Consortium
1. Laboratorio de Microbiologia Molecular y Estructural, Institut Pasteur de Montevideo, Montevideo, Uruguay
2. Unidad Mixta UMPI, Institut Pasteur de Montevideo + Instituto Nacional de Investigacion Agropecuaria INIA, Montevideo, Uruguay
3. Departamento de Bacteriologia, Division Laboratorios Veterinarios "Miguel C. Rubino" Sede Central, Ministerio de Ganaderia, Agricultura y Pesca, Montevideo, Uruguay
4. Departamento de Bacteriologia y Virologia, Instituto de Higiene, Facultad de Medicina, Universidad de la Republica, Montevideo, Uruguay
5. Division Laboratorios Veterinarios "Miguel C. Rubino" Laboratorio Regional Noroeste, Ministerio de Ganaderia, Agricultura y Pesca, Paysandu, Uruguay
6. Division Laboratorios Veterinarios "Miguel C. Rubino" Laboratorio Regional Este, Ministerio de Ganaderia, Agricultura y Pesca, Treinta y Tres, Uruguay
7. Instituto Nacional de Investigacion Agropecuaria INIA, Estacion Experimental La Estanzuela, Colonia, Uruguay
8. Unité de Biologie des Spirochètes, Institut Pasteur, Paris, France,
9. Joint International Unit "Integrative Microbiology of Zoonotic Agents" IMiZA, Institut Pasteur de Montevideo, Montevideo, Uruguay / Institut Pasteur, Paris, France
10. Département de Microbiologie, Institut Pasteur, Paris, France
* These authors contributed equally to this work.
** These authors also contributed equally to this work