Describing the genetic diversity of human populations is essential to improve our understanding of human diseases and their geographical distribution. However, the vast majority of genetic studies have been focused on populations of European ancestry, which represent only 16% of the global population. Scientists at the Institut Pasteur, Collège de France, and CNRS have looked at understudied human populations from the South Pacific, which are severely affected by a variety of diseases, including vector-borne infectious diseases such as Zika virus, dengue, and chikungunya, and metabolic diseases such as obesity and diabetes. Using genome sequencing of 320 individuals, the scientists have investigated how human populations have biologically adapted to the environments of the Pacific islands and how this has affected their current state of health. This study has also revealed hitherto unsuspected aspects of the history of human settlement in this region. This work is published in the April 14th, 2021 issue of Nature.
An international consortium of scientists organized by Etienne Patin (CNRS/Institut Pasteur) and Lluis Quintana-Murci (Collège de France/Institut Pasteur) was set up to characterize the genetic diversity of populations in the South Pacific, a region full of contrasts with its myriad islands that have been settled at very different time periods.
Indeed, shortly after humans left Africa, they settled Near Oceania (Papua-New-Guinea, the Bismarck Archipelago, and the Solomon Islands) approximately 45,000 years ago, while the rest of the Pacific, known as Remote Oceania (Vanuatu, the Wallis and Futuna Islands, Polynesia, etc.), remained uninhabited. It was only approximately 40,000 years later that Remote Oceania was peopled: it is currently accepted that a group of humans left Taiwan 5,000 years ago - a migration known as the ‘Austronesian expansion’ - passed through the Philippines, Indonesia, and the already-inhabited Near Oceanian islands to eventually settle Remote Oceania for the first time.
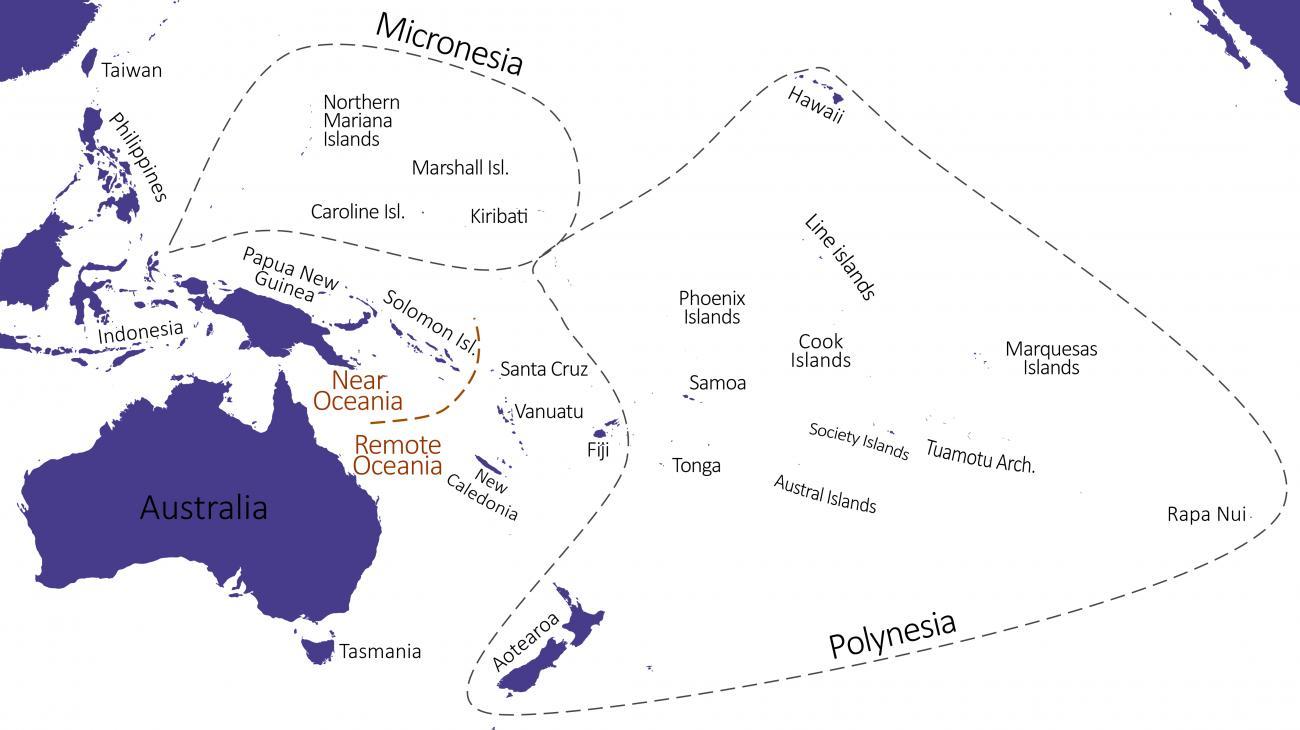
Credit : Institut Pasteur
En route to these remote lands, the ancestors of South Pacific populations met with groups of archaic humans, with whom they interbred. While 2-3% of modern-day Oceanian populations’ genetic material is inherited from Neanderthals (which all populations outside Africa also possess), up to 3% of their genomes is also inherited from Denisovans (relatives of Neanderthals thought to have originated in Asia). It was already known that modern humans inherited beneficial mutations from Neanderthals via admixture, which improved their ability to adapt to their environment, including resistance to viral infections (Cell, 20161). In this study published today, scientists from the Human Evolutionary Genetics Unit (Institut Pasteur/CNRS2) in collaboration with various laboratories in France3, Germany, Sweden, Switzerland, China, and Taiwan have sought to shed light on how ancient admixture helped Pacific populations to adapt to their specific island environments, including any pathogens encountered.
Historical events traced using genetics
Based on whole-genome-sequencing of over 320 individuals from Taiwan, the Philippines, the Bismarck Archipelago, the Solomon Islands, Santa Cruz Islands, and Vanuatu, this work published in the Nature journal has helped tracing the history of the human settlement of Oceania. First, the scientists have dated the settlement of the various islands of Near Oceania back to approximately 40,000 years, thus confirming archaeological records. They have also demonstrated that this initial settlement was followed by a period of genetic isolation between islands. "Our results confirm that humans were able to cross the seas to reach new lands from an early stage. However, they also suggest that these voyages were relatively infrequent at this distant period in history," explains Etienne Patin, a CNRS scientist within the Human Evolutionary Genetics Unit at the Institut Pasteur. Moreover, the results of the study reveal a major reduction in the size of these populations just before the settlement.
Second, the study challenges the theory, known as the Out-of-Taiwan model, that a population left Taiwan approximately 5,000 years ago to rapidly settle Near and Remote Oceania. "Our analyses suggest that humans left Taiwan more than 5,000 years ago, and that admixture between the Austronesian incomers and the populations of Near Oceania started only 2,000 years later. The expansions from Taiwan therefore took some time, and may have involved a maturation phase in the Philippines or Indonesia," comments Etienne Patin.
The heritage of Neanderthals and Denisovans in South Pacific populations
Through this work, it has been possible to estimate the proportions of Neanderthal and Denisovan material in the genomes of South Pacific populations. "We were surprised to note that, contrary to the Neanderthal heritage, which is very similar among the twenty populations studied (approximately 2.5%), the Denisovan legacy varies considerably between populations, from virtually 0% in Taiwan and the Philippines to up to 3.2% in Papua-New-Guinea and Vanuatu" comments Lluis Quintana-Murci, Professor at the Collège de France, holder of the Chair in Human Genomics and Evolution, and Head of the Human Evolutionary Genetics Unit (Institut Pasteur/CNRS).
But this was not the only surprise. The study confirms that Neanderthals provided modern human populations with beneficial mutations associated with numerous phenotypes: skin pigmentation, metabolism, neuronal development, etc. However, the most surprising finding is that admixture with Denisovans has almost exclusively brought beneficial mutations related to immune response regulation. This suggests that the Denisovan heritage has been a reservoir of advantageous mutations, which have improved the Pacific populations' ability to survive local pathogens. Therefore, it appears that these populations have benefited from the advantages bestowed through admixture with both forms of archaic humans.
Furthermore, the study demonstrates that admixture with Denisovans did not occur at once, but over the course of at least four independent events. It shows that the Denisovans with whom the Pacific populations interbred were in fact highly diverse populations. This conclusion was impossible to deduce from the single genome of a Denisovan specimen found in Siberia: "One of the strengths of these analyses is that, by studying the 3% of archaic heritage present in the genomes of modern humans, one can ‘resurrect’ Denisovans' genomes, and thus show that they presented high levels of genetic diversity," comments Lluis Quintana-Murci.
Finally, besides the biological adaptation enabled by archaic admixture, the scientists have found that lipid metabolism, cholesterol in particular, was also a target of natural selection among Oceanian peoples. This insight will potentially improve our understanding of why recent changes in these populations' lifestyle may be associated with metabolic disorders.
By taking an evolutionary genetics approach, it is possible to shed light on the history of populations' biological adaptation to their environment, and provide the scientific community with valuable information on specific human traits. These large-scale genomic studies will eventually help us better understand the genetic causes of diseases affecting some regions of the globe, where medical research has hitherto been scarce.
1 Genetic Adaptation and Neandertal Admixture Shaped the Immune System of Human Populations, Cell, 20 octobre 2016.
2 Evolutionary genomics, modeling and health (GEMS)
3 The following were among the laboratories involved: Virology (CNRS/Institut Pasteur), Eco-anthropology (EAE, CNRS/MNHN/Université de Paris), and Archeology and Sciences of Antiquity (ArScAn, CNRS/Université Paris 1 Panthéon-Sorbonne/Université Paris Nanterre/Ministry of Culture)
Source
Genomic insights into population history and biological adaptation in Oceania, Nature, April 14th, 2021
Jeremy Choin1,2,15, Javier Mendoza-Revilla1,15, Lara R. Arauna1,15, Sebastian Cuadros-Espinoza1,3, Olivier Cassar4, Maximilian Larena5, Albert Min-Shan Ko6, Christine Harmant1, Romain Laurent7, Paul Verdu7, Guillaume Laval1, Anne Boland8, Robert Olaso8, Jean-François Deleuze8, Frédérique Valentin9, Ying-Chin Ko10, Mattias Jakobsson5, Antoine Gessain4, Laurent Excoffier11,12, Mark Stoneking13, Etienne Patin1,16* & Lluis Quintana-Murci1,14,16
1 Human Evolutionary Genetics Unit, Institut Pasteur, UMR 2000, CNRS, Paris 75015, France
2 Université Paris Diderot, Sorbonne Paris Cité, Paris 75013, France
3 Sorbonne Université, Collège doctoral, Paris 75005, France
4 Oncogenic Virus Epidemiology and Pathophysiology, Institut Pasteur, UMR 3569, CNRS, Paris 75015, France
5 Human Evolution, Department of Organismal Biology and SciLife lab, Uppsala University, Uppsala 752 36, Sweden
6 Key Laboratory of Vertebrate Evolution and Human Origins, Institute of Vertebrate Paleontology and Paleoanthropology, Chinese Academy of Sciences, Beijing 100044, China
7 Muséum National d’Histoire Naturelle, UMR7206, CNRS, Université de Paris, Paris 75016, France
8 Centre National de Recherche en Génomique Humaine (CNRGH), Institut de Biologie François Jacob, CEA, Université Paris-Saclay, Evry 91057, France
9 Maison de l’Archéologie et de l’Ethnologie, UMR 7041, CNRS, Nanterre 92023, France
10 Environment-Omics-Disease Research Center, China Medical University and Hospital, Taichung 40447, Taiwan 11Institute of Ecology and Evolution, University of Bern, Bern 3012, Switzerland
12 Swiss Institute of Bioinformatics, Lausanne 1015, Switzerland
13 Department of Evolutionary Genetics, Max Planck Institute for Evolutionary Anthropology, Leipzig 04103, Germany
14 Chair Human Genomics and Evolution, Collège de France, Paris 75005, France
15 These authors contributed equally
16 These authors jointly supervised this work


